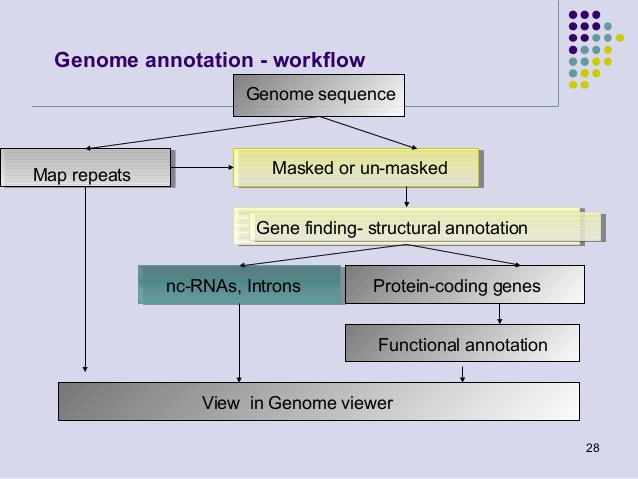
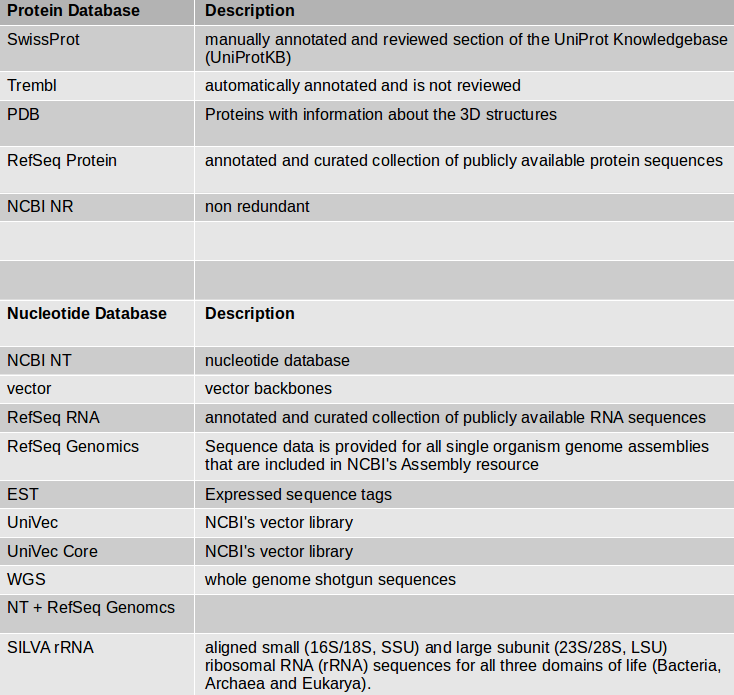
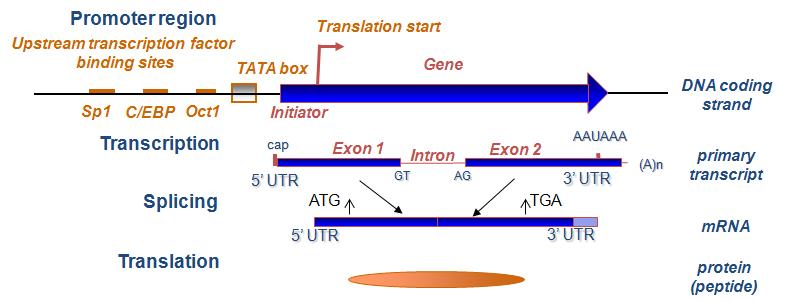
Comparison of different annotation tools for characterization of the complete chloroplast genome of Corylus avellana cv Tombul | BMC Genomics | Full Text
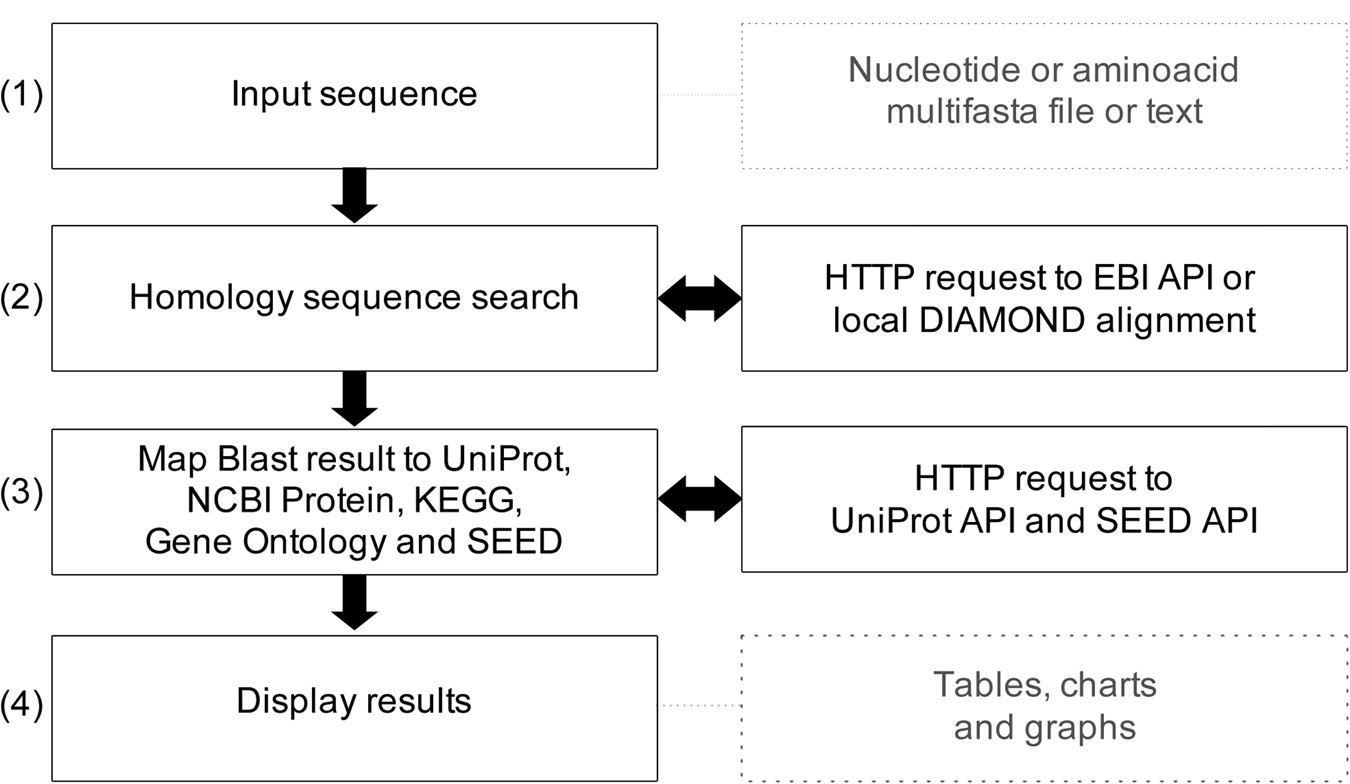
GO FEAT: a rapid web-based functional annotation tool for genomic and transcriptomic data | Scientific Reports

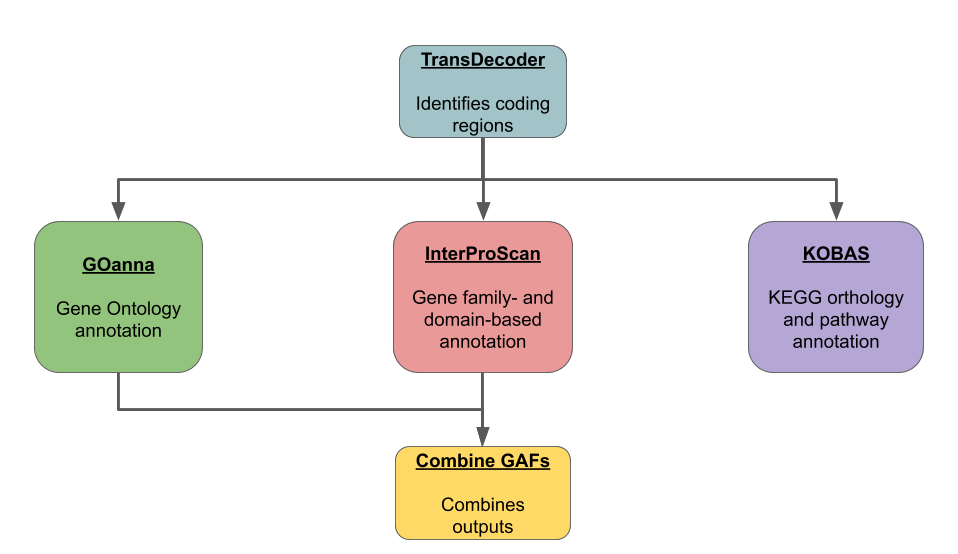
GitHub - galikbence/genome_annotation: Pipeline for eukaryotic genome annotation based on external evidences

BRAKER2: Automatic Eukaryotic Genome Annotation with GeneMark-EP+ and AUGUSTUS Supported by a Protein Database | bioRxiv

Genome Research on X: "@genomeresearch publishes a Perspective focusing on the expression of alternative ORFs, the current tools for studying them, and proposes a genome annotation framework that includes alternative ORFs to